
Geographic distance
distance.RdIf x is a SpatRaster:
If y is missing this method computes the distance, for all cells that are NA in SpatRaster x to the nearest cell that is not NA (or other values, see arguments "target" and "exclude").
If y is a numeric value, the cells with that value are ignored. That is, distance to or from these cells is not computed.
If y is a SpatVector, the distance to that SpatVector is computed for all cells, optionally after rasterization.
The distance is always expressed in meter if the coordinate reference system is longitude/latitude, and in map units otherwise. Map units are typically meter, but inspect crs(x) if in doubt.
Results are more precise, sometimes much more precise, when using longitude/latitude rather than a planar coordinate reference system, as these distort distance.
If x is a SpatVector:
If y is missing, a distance matrix between all objects in x is computed. A distance matrix object of class "dist" is returned.
If y is a SpatVector, the geographic distance between all objects is computed (and a matrix is returned). If both sets have the same number of points, and pairwise=TRUE, the distance between each pair of objects is computed, and a vector is returned.
If x is a matrix:
x should consist of two columns, the first with "x" (or longitude) and the second with "y" coordinates (or latitude). If y is a also a matrix, the distance between each point in x and all points in y is computed, unless pairwise=TRUE
If y is missing, the distance between each point in x with all other points in x is computed, unless sequential=TRUE
Usage
# S4 method for class 'SpatRaster,missing'
distance(x, y, target=NA, exclude=NULL, unit="m", method="haversine",
maxdist=NA, values=FALSE, filename="", ...)
# S4 method for class 'SpatRaster,SpatVector'
distance(x, y, unit="m", rasterize=FALSE, method="haversine", filename="", ...)
# S4 method for class 'SpatVector,SpatVector'
distance(x, y, pairwise=FALSE, unit="m", method="haversine",
use_nodes=FALSE, names=NULL)
# S4 method for class 'SpatVector,ANY'
distance(x, y, sequential=FALSE, pairs=FALSE, symmetrical=TRUE, unit="m",
method="haversine", use_nodes=FALSE, names=NULL)
# S4 method for class 'matrix,matrix'
distance(x, y, lonlat, pairwise=FALSE, unit="m", method="geo")
# S4 method for class 'matrix,missing'
distance(x, y, lonlat, sequential=FALSE, pairs=FALSE, symmetrical=TRUE,
unit="m", method="geo")Arguments
- x
SpatRaster, SpatVector, or two-column matrix with coordinates (x,y or lon,lat)
- y
missing, numeric, SpatVector, or two-column matrix
- target
numeric. The value of the cells for which distances to cells that are not
NAshould be computed- exclude
numeric. The value of the cells that should not be considered for computing distances
- unit
character. Can be either "m" or "km"
- method
character. One of "geo", "cosine" or "haversine". With "geo" the most precise but slower method of Karney (2003) is used. The other two methods are faster but less precise
- maxdist
numeric. Distances above this value are set to
NA- values
logical. If
TRUE, the value of the nearest non-target cell is returned instead of the distance to that cell- rasterize
logical. If
TRUEdistance is computed from the cells covered by the geometries after rasterization. This can be much faster in some cases- filename
character. Output filename
- ...
additional arguments for writing files as in
writeRaster- sequential
logical. If
TRUE, the distance between sequential geometries is returned- pairwise
logical. If
TRUEand if x and y have the same size (number of rows), the pairwise distances are returned instead of the distances between all elements- lonlat
logical. If
TRUEthe coordinates are interpreted as angular (longitude/latitude). IfFALSEthey are interpreted as planar- pairs
logical. If
TRUEa "from", "to", "distance" matrix is returned- symmetrical
logical. If
TRUEandpairs=TRUE, the distance between a pair is only included once. The distance between geometry 1 and 3 is included, but the (same) distance between 3 and 1 is not- use_nodes
logical. If
TRUEand the crs is longitude/latitude, the nodes (vertices) of lines or polygons are used to compute distances, instead of the lines that connect them. This is faster, but can be less precise if the nodes are far apart- names
character. One (or two) variable names in
x(andy) to label the distance matrix
References
Karney, C.F.F., 2013. Algorithms for geodesics, J. Geodesy 87: 43-55. doi:10.1007/s00190-012-0578-z.
Examples
#lonlat
r <- rast(ncols=36, nrows=18, crs="+proj=longlat +datum=WGS84")
r[500] <- 1
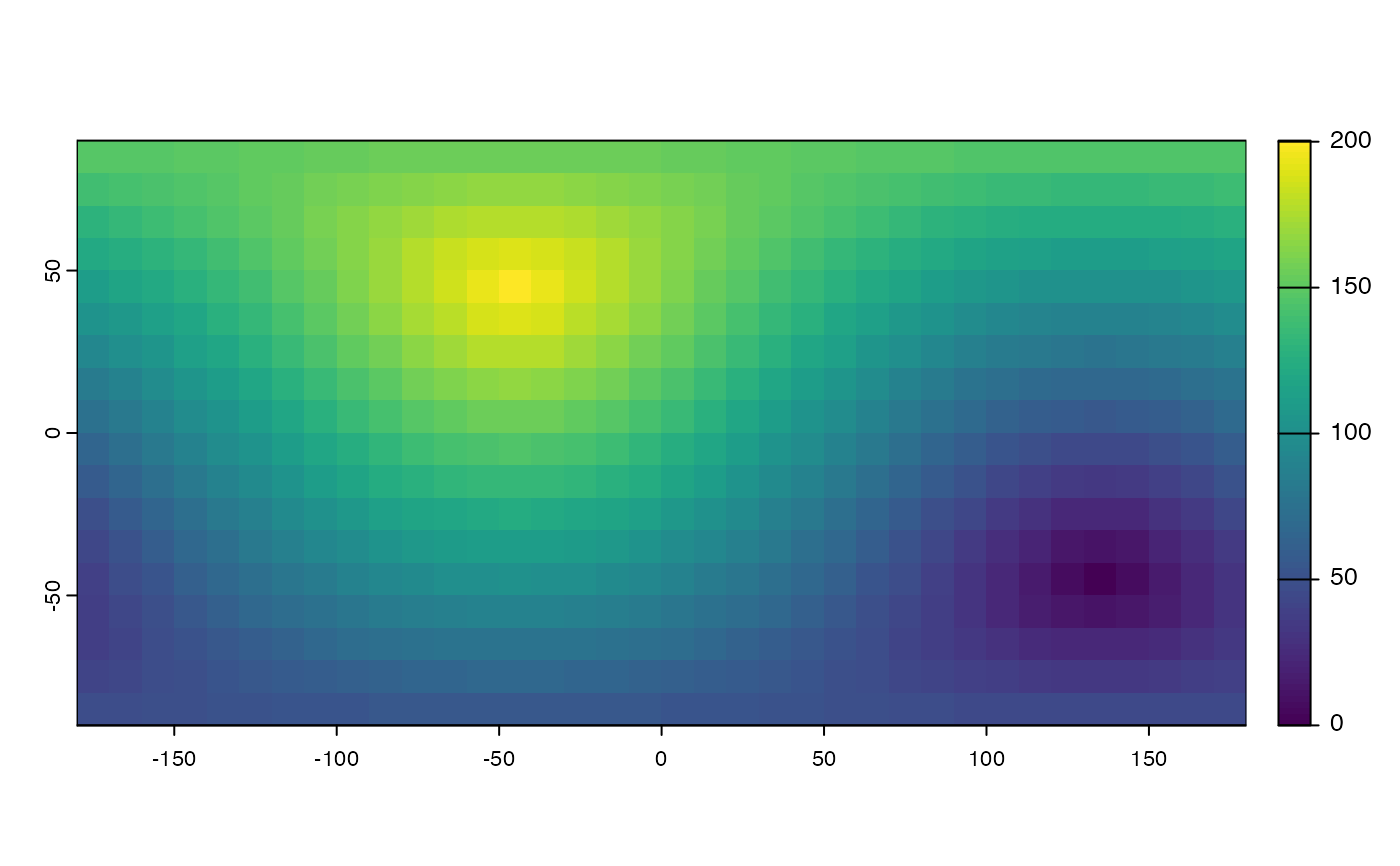
d <- distance(r, unit="km")
plot(d / 1000)
 #planar
rr <- rast(ncols=36, nrows=18, crs="+proj=utm +zone=1 +datum=WGS84")
rr[500] <- 1
d <- distance(rr)
rr[3:10, 3:10] <- 99
e <- distance(rr, exclude=99)
p1 <- vect(rbind(c(0,0), c(90,30), c(-90,-30)), crs="lonlat")
values(p1) <- data.frame(ID=LETTERS[1:3])
dp <- distance(r, p1)
d <- distance(p1)
d
#> 1 2
#> 2 10018754
#> 3 10018754 20037508
as.matrix(d)
#> 1 2 3
#> 1 0 10018754 10018754
#> 2 10018754 0 20037508
#> 3 10018754 20037508 0
p2 <- vect(rbind(c(30,-30), c(25,40), c(-9,-3)), crs="+proj=longlat +datum=WGS84")
values(p2) <- data.frame(ID=letters[1:3])
dd <- distance(p1, p2, names=c("ID", "ID"))
dd
#> a b c
#> A 4609698 5124121 1055634
#> B 9219396 5900335 11053086
#> C 10818112 14137173 8984422
pd <- distance(p1, p2, pairwise=TRUE)
pd
#> [1] 4609698 5900335 8984422
pd == diag(dd)
#> [1] TRUE TRUE TRUE
# polygons, lines
crs <- "+proj=utm +zone=1"
p1 <- vect("POLYGON ((0 0, 8 0, 8 9, 0 9, 0 0))", crs=crs)
p2 <- vect("POLYGON ((5 6, 15 6, 15 15, 5 15, 5 6))", crs=crs)
p3 <- vect("POLYGON ((2 12, 3 12, 3 13, 2 13, 2 12))", crs=crs)
p <- rbind(p1, p2, p3)
L1 <- vect("LINESTRING(1 11, 4 6, 10 6)", crs=crs)
L2 <- vect("LINESTRING(8 14, 12 10)", crs=crs)
L3 <- vect("LINESTRING(1 8, 12 14)", crs=crs)
lns <- rbind(L1, L2, L3)
pts <- vect(cbind(c(7,10,10), c(3,5,6)), crs=crs)
distance(p1,p3)
#> [,1]
#> [1,] 3
distance(p)
#> 1 2
#> 2 0
#> 3 3 2
distance(p,pts)
#> [,1] [,2] [,3]
#> [1,] 0.000000 2.000000 2.000000
#> [2,] 3.000000 1.000000 0.000000
#> [3,] 9.848858 9.899495 9.219544
distance(p,lns)
#> [,1] [,2] [,3]
#> [1,] 0.000000 3.535534 0.000000
#> [2,] 0.000000 0.000000 0.000000
#> [3,] 1.414214 5.099020 2.553878
distance(pts,lns)
#> [,1] [,2] [,3]
#> [1,] 3 8.602325 7.262591
#> [2,] 1 5.385165 6.943356
#> [3,] 0 4.472136 6.065460
#planar
rr <- rast(ncols=36, nrows=18, crs="+proj=utm +zone=1 +datum=WGS84")
rr[500] <- 1
d <- distance(rr)
rr[3:10, 3:10] <- 99
e <- distance(rr, exclude=99)
p1 <- vect(rbind(c(0,0), c(90,30), c(-90,-30)), crs="lonlat")
values(p1) <- data.frame(ID=LETTERS[1:3])
dp <- distance(r, p1)
d <- distance(p1)
d
#> 1 2
#> 2 10018754
#> 3 10018754 20037508
as.matrix(d)
#> 1 2 3
#> 1 0 10018754 10018754
#> 2 10018754 0 20037508
#> 3 10018754 20037508 0
p2 <- vect(rbind(c(30,-30), c(25,40), c(-9,-3)), crs="+proj=longlat +datum=WGS84")
values(p2) <- data.frame(ID=letters[1:3])
dd <- distance(p1, p2, names=c("ID", "ID"))
dd
#> a b c
#> A 4609698 5124121 1055634
#> B 9219396 5900335 11053086
#> C 10818112 14137173 8984422
pd <- distance(p1, p2, pairwise=TRUE)
pd
#> [1] 4609698 5900335 8984422
pd == diag(dd)
#> [1] TRUE TRUE TRUE
# polygons, lines
crs <- "+proj=utm +zone=1"
p1 <- vect("POLYGON ((0 0, 8 0, 8 9, 0 9, 0 0))", crs=crs)
p2 <- vect("POLYGON ((5 6, 15 6, 15 15, 5 15, 5 6))", crs=crs)
p3 <- vect("POLYGON ((2 12, 3 12, 3 13, 2 13, 2 12))", crs=crs)
p <- rbind(p1, p2, p3)
L1 <- vect("LINESTRING(1 11, 4 6, 10 6)", crs=crs)
L2 <- vect("LINESTRING(8 14, 12 10)", crs=crs)
L3 <- vect("LINESTRING(1 8, 12 14)", crs=crs)
lns <- rbind(L1, L2, L3)
pts <- vect(cbind(c(7,10,10), c(3,5,6)), crs=crs)
distance(p1,p3)
#> [,1]
#> [1,] 3
distance(p)
#> 1 2
#> 2 0
#> 3 3 2
distance(p,pts)
#> [,1] [,2] [,3]
#> [1,] 0.000000 2.000000 2.000000
#> [2,] 3.000000 1.000000 0.000000
#> [3,] 9.848858 9.899495 9.219544
distance(p,lns)
#> [,1] [,2] [,3]
#> [1,] 0.000000 3.535534 0.000000
#> [2,] 0.000000 0.000000 0.000000
#> [3,] 1.414214 5.099020 2.553878
distance(pts,lns)
#> [,1] [,2] [,3]
#> [1,] 3 8.602325 7.262591
#> [2,] 1 5.385165 6.943356
#> [3,] 0 4.472136 6.065460